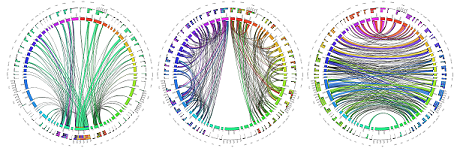
Lifespan Developing Human Connectome Project (dHCP) Study
The dHCP study planned to enroll 1500 Subjects at age 20-44 weeks post-conception. The purpose is to link together imaging, clinical, behavioural, and genetic information..
License
The derived files below are shared under the dHCP data sharing agreement. The source of the data are from the 3rd release.
Users using the files should follow agreement and cite/acknowledge the source.
Download
Methods
A multishell diffusion scheme was used, and the b-values were 400 ,1000 and 2600 s/mm². The number of diffusion sampling directions were 64, 88, and 128, respectively. The in-plane resolution was 1.5 mm. The slice thickness was 1.5 mm. The images were denoised and corrected for Gibbs ringing, motion, eddy current, and susceptibility artifact using the diffusion SHARD pipeline. A quality check was conducted using neighboring DWI correcltion (NDC) (Yeh, Neuroimage. 2019 Nov 15;202:116131). 34 out of 758 scans (including repeated scans) were excluded due to their low NDC values identified by a median value based outlier detector. The accuracy of b-table orientation was examined by comparing fiber orientations with those of a population-averaged template (Yeh et al. Neuroimage, 2018). The restricted diffusion was quantified using restricted diffusion imaging (Yeh et al., MRM, 77:603–612 (2017)). The diffusion data were reconstructed using generalized q-sampling imaging (Yeh et al., IEEE TMI, ;29(9):1626-35, 2010) with a diffusion sampling length ratio of 1.25. The tensor metrics were calculated. The analysis was conducted using the resource allocation (TG-CIS200026) at Extreme Science and Engineering Discovery Environment (XSEDE) resources (Towns, J. et al. Computing in science & engineering 16, 62-74 2014).
This analysis used Bridges-2 at Pittsburgh Supercomputer Center through allocation CIS200026 from the Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS) program (Boerner TJ, Deems S, Furlani TR, Knuth SL, Towns J. Access: Advancing innovation: Nsf’s advanced cyberinfrastructure coordination ecosystem: Services & support. InPractice and Experience in Advanced Research Computing 2023 Jul 23 pp. 173-176).
- 642 FIB files (Ready-to-track using DSI Studio)
- 642 SRC files (none repeated scans, including twins)
- 164 Scan-Rescan SRC files (n=82) Repeated scan of the same subjects. All subjects are also included in the 642 SRC files above. These SRC files are preprocessed results from the diffusion SHARD pipeline. The background signals are zeroed and image dimension cropped. The data are largely reduced and CANNOT be reversed back to the original data distribution.
- Connectometry DB for correlational or differential tractography
- Demographics
Processing Steps
1. generate SRC files from NIFTI files
Copy all NIFTI (DWI and mask), bval, bvec files to the same folder and use DSI Studio’s GUI Batch function [Batch Processing[Step B2b: NIFTI to SRC (Single Folder)]
2. reconstruction
This was done using DSI Studio GUI. Click on [Step T2 Reconstruction] and select all SRC files.
- [Step T2a][Edit][Open] to load mask
- [Edit][Crop Background]
- [Run Reconstruction]
3. fiber Tracking
The following script for a job arrary to runs fiber tracking on all FIB file.
#!/bin/bash
subs=$(ls -L *.fib.gz)
subs=(${subs// /})
echo "processing ${subs[$1]}"
singularity exec dsistudio_latest.sif dsi_studio --action=atk --export_template_trk=1 --source=${subs[$1]}
The following is the job array to run the above script using sbatch. The script needs an the file name of the script to run the job array.
#!/bin/bash
#SBATCH -t 24:00:00
#SBATCH -p RM-shared
#SBATCH -N 1
#SBATCH --ntasks-per-node 8
#SBATCH --mem=15GB
#SBATCH --array=0-999
set -x
sh $1 $SLURM_ARRAY_TASK_ID