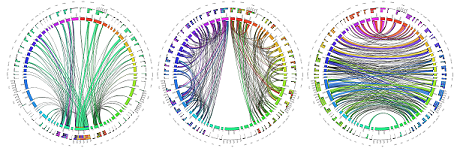
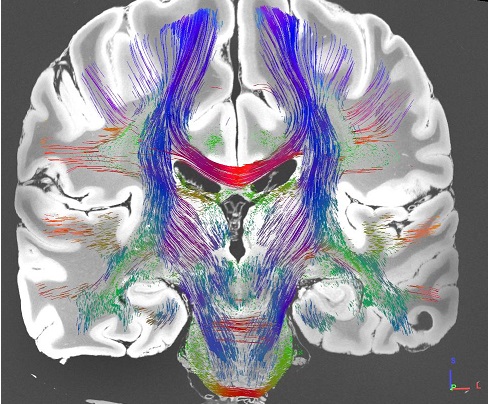
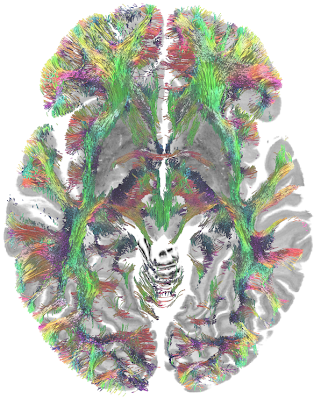
MGH Single Subject 760µm dMRI data

Download
-
The FIB file is ready-to-track and can be opened in DSI Studio to perform fiber tracking. To track the data, download DSI Studio at dsi-studio.labsolver.org and open the FIB.gz file in [Step T3 Fiber Tracking]. The figures were created by whole brain tracking of 500,000 tracks and cut at mid-brain and pons level using [Edit][Cut by slice][Cut Z+]
The SRC file contains raw dMRI signals and can be used to generate FIB files. The TT file under /tract folder are tractography files.
- sub-01_dwi.qsdr.fz (DWI data reconstructed to the ICBM152 space, ready to align with BigBrain and other ICBM152 space data)
- sub-01_dwi.gqi.fz (DWI data reconstructed in the native space)
- sub-01_dwi.sz (DWI signals and associated b-table aggregated in one file)
Methods
A multishell diffusion scheme was used, and the b-values were 1000 and 2500 s/mm2. The number of diffusion sampling directions were 840 and 1680, respectively. The in-plane resolution was 0.76 mm. The slice thickness was 0.76 mm. The diffusion data were reconstructed in the MNI space at 0.5 mm using q-space diffeomorphic reconstruction (Yeh et al., Neuroimage, 58(1):91-9, 2011) to obtain the spin distribution function (Yeh et al., IEEE TMI, ;29(9):1626-35, 2010). A diffusion sampling length ratio of 1.25 was used The output resolution of is 0.5 mm isotorpic. The restricted diffusion was quantified using restricted diffusion imaging (Yeh et al., MRM, 77:603–612 (2017)).
License
- The source of the data was shared under CC0 license at 1 and 2, and I would strongly recopmmend acknowledging the data source to the reference (Wang et al at MGH)
- The SRC and FIB data shared on this page were processed by DSI Studio and inhereted the same CC0 license. Please acknowledge Wang et al. at MGH and mentioned the data were processed in DSI Studio.
Reference
Wang F, Dong Z, Tian Q, Liao C, Fan Q, Hoge WS, Keil B, Polimeni JR, Wald LL, Huang SY, Setsompop K. In vivo human whole-brain Connectom diffusion MRI dataset at 760 µm isotropic resolution. Scientific Data. 2021 Apr 29;8(1):1-2.
MGH Single Subject 100µm MRI
Download
-
The NIFTI file is an image volume. It is mainly used to reveal detailed structures and cannot be used to track fiber pathways.
Reference
Edlow, B.L., Mareyam, A., Horn, A., Polimeni, J.R., Witzel, T., Tisdall, M.D., Augustinack, J.C., Stockmann, J.P., Diamond, B.R., Stevens, A. and Tirrell, L.S., 2019. 7 Tesla MRI of the ex vivo human brain at 100 micron resolution. Scientific data, 6(1), pp.1-10.